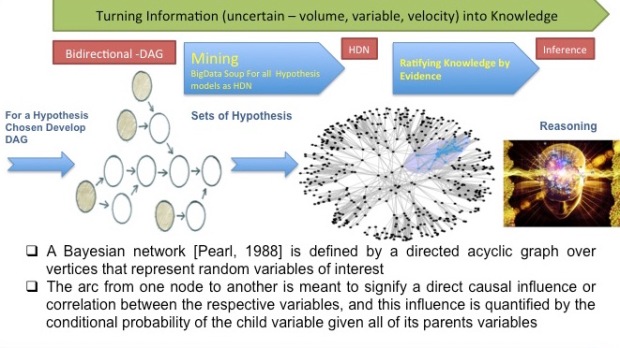
METHODS USED IN The BioIngine APPROACH: ROOTS OF THE HYPERBOLIC DIRAC NETWORK (HDN). – Dr. Barry Robson
General Approach : Solving the Representation and Use of Knowledge for the Real World.
Blending Systematically Produced and Unsystematically Existing Information and Synthesizing the Knowledge.
The area of our efforts in the support of healthcare and biomedicine is essentially one in Artificial Intelligence (AI). For us, however, this means a semantic knowledge engineering approach intimately combined with principles of probability theory, information theory, number theory, theoretical physics, data analytic principles, and even linguistic theory. These contributions and the unification of these, in the manner described briefly later below, is the general theory of an entity called the Hyperbolic Dirac Net (HDN), a means of representing and probabilistically quantifying networks of knowledge of both a simple probabilistic, and an even more sophisticated probabilistic semantic, nature in a way that has not been possible for previous approaches. It provides the core methodology for making use of medical knowledge in the face of considerable uncertainty and risk in the practice of medicine, and not least the need to manage massive amounts of diverse data, including both structured data and unstructured natural language text. As described here, the ability of the HDN and its supporting Q-UEL language to handle also the kind of interactions between things that we describe in natural language by using verbs and propositions, take account of the complex lacework of interactions between things, and do so when our knowledge is of probabilistic character, are of pressing and crucial importance to development of a higher level of information technology in many fields, but particularly in medicine.
In a single unified strike, the mathematics of the HDN, adapted in a virtually seamlessand natural way from a standard in physics due to Nobel Laureate Paul Dirac as discussed below, addresses several deficiencies (both well-known and less well advertised) in current forms of automated inference. These deficiencies largely relate to assumptions and representations that are not fully representative of the real world. They are touched upon later below, but the general one of most strategic force is as follows. As is emphasized and as discussed here, of essential importance to modern developments in many industries and disciplines, and not least in medicine, is the capture of large amounts of knowledge in what we call a Knowledge Representation Store (KRS). Each entry or element in such a store is a statement about the world. Whatever the name, the captured knowledge includes basic facts and definitions about the world in general, but also knowledge about specific cases (and looking more like what is often meant by “data”), such as a record about the medical status of a patient or a population. From such a repository of knowledge, general and specific, end users can invoke automated reasoning and inference to predict, aid decision making, and move forward acting on current best evidence Wide acceptance and pressing need is demonstrated (see below) by numerous efforts from the earliest Expert systems to the emerging Semantic Web, an international effort to link not just web pages (as with the World Wide Web) but also data and knowledge, and comparable efforts such as Never-Ending Language Learning system (NELL) at Carnegie Mellon University. The problem is that there is no single agreed way to actually using such a knowledge store in automated reasoning and inference, especially when uncertainty is involved.
In part this problem is perhaps in part because there is the sense that there is something deep that is still missing in what we mean by “Artificial Intelligence” (AI), and in part by lack of agreement in how to reason with connections of knowledge represented as a general graph. The latter is even to the extent that the popular Bayes Net is, by its original definition, a directed acyclic graph (DAG) that ignores or denies cyclic paths in knowledge networks, in stark contrast to the multiple interactions in a “mind map” concept map in student study notes, a subway map, biochemical pathways, physiological interactions, the wiring of the human brain, and the network of interactions in ecology. Primarily, however, the difficulty is that the elements of knowledge in the Semantic Web and other KRS-like efforts are for the most part presented as authoritative assertions rather than treated probabilistically. This is the despite the fact that the pioneering Expert Systems for medicine needed from the outset to be essentially probabilistic in order to manage uncertainty in the knowledge used to make decisions and the combining of it, and to deduce most probable diagnosis and select best therapy amongst many initial options, although here too there is lack of agreement, and almost every new method represented a different perception and use of uncertainty. Many of the aspects, use of a deeper theory, arrangement of knowledge elements into a general graph, might be addressed in the way a standard repository of knowledge is used, i.e. applied after a KRS is formed, but a proper and efficient treatment can only associate probability with the elements of represented knowledge from the outset (even though, like any aspect of knowledge, the probabilities should be allowed to evolve by refinement and updating). One cannot apply a probabilistic logic without probabilities in the axioms, or at least not to any advantage. Further, it makes no sense to have elements of knowledge, however they are used, that state unequivocally that some things are true, e.g. that obese patients are type 2 diabetics, because it is a matter of probability, in this case describing the scope of applicability of the statement to patients, i.e. only some 20-30% are so. Indeed, in that case, using only certainty or near-certainty, this medically significant association might never have appeared as a statement in the first place. Note that the importance of probabilistic thinking is also exemplified here by the fact that the reader may have been expecting or thinking in terms of “type 2 patients are obese”, which is not the same thing and has a probability of about 90%, closer to certainty, but noticeably still not 100%. All the above aspects, including the latter “two way” diabetes example, relate to matters that are directly relevant, and the differentiating features, of an HDN. The world that humans perceive is full of interactions in all directions, yet full of uncertainty, so we cannot only say that
“HDN = Semantic Knowledge + General Graph + Probability = Best Decision Making”
but also that any alternative method runs the risk of being seriously wrong or severely approximate if ignores any of knowledge or general graph or probability. For example, the popular Bayes Net as discussed below is probabilistic, but it uses only conditional and prior probabilities as knowledge, is a very restricted form of graph. Conversely, approach like that of IBM’s well-known Watson is clearly limited, and leaves a great deal to be sifted, corrected, and reasoned by the user, if is primarily a matter of “a super search engine” rather than inferring from an intricate lacework of probabilistic interactions. Importantly, even if it might be argued that some areas of science and industry can for the most part avoid such subtleties relating to probability, it is certainly not true in medicine, as the above diabetes example illustrates. From the earliest days of clinical decision support it clearly made no sense to pick, for example, “a most true diagnosis” from a set of possible diagnoses each registered only, on the evidence available so far, as true or false. What is vitally important to medicine is a semantic system that the real world merits, one capable of handling degree of truth and uncertainty in a quantitative way. Our larger approach, additionally building on semantic and linguistic theory, can reasonably be called probabilistic semantics. By knowledge in an HDN we also mean semantic knowledge in general, including that expressed by statements with relationships that are verbs of actions. In order to be able also to draw upon the preexisting Semantic Web and other efforts that contain such statements, however, the HDN approach is capable of making use of knowledge represented as certain[2].
Knowledge and reasoning from it does not stand alone from the rest of information management in the domain that generates and uses it, and it is a matter to be seriously attended to when, in comparison to many other industries such as finance, interoperability and universally accepted standards are lacking. Importantly, the application of our approach, and our strategy for healthcare and biomedicine, covers a variety of areas in healthcare information technology that we have addressed as proofs-of-concept in software development, welded into a single focus by a unification made possible through the above theoretical and methodological principles. These areas include digital patient records, privacy and consent mechanisms, clinical decision support, and translational research (i.e. getting the results of relevant biomedical research such as new genomics findings to physicians faster). All of these are obviously required to provide information for actions taken by physicians and other medical workers, but the broad sweep is also essential because no aspect stands alone: there has been a need for new semantic principles, based on the core features of the AI approach, to achieve interoperability and universal exchange.
- There are various terms for such a knowledge store. “Knowledge Representation Store” is actually our term emphasizing that it is (in our view) analogous to human memory as enabled and utilized by human thought and language, but now in a representation that computers can readily read directly and use efficiently (while in our case also remaining readable directly by humans in a natural way).
- In such cases, probability one (P=1) is the obvious assignment, but strictly speaking in our approach this technically means that it is an assertion that awaits refutation, in the manner of the philosophy of Karl Popper, and consistent with information theory in which the information content I of any statement of probability P is I = -ln(P), i.e. we find information I=0 when probability P=1. A definition such as “cats are mammals” seems an exception, but then, as long as it stands as a definition, it will not be refuted.
- These are the rise of medical IT (and AI in general) as the next “Toffler wave of industry”, the urgent need to greatly reduce inefficiency and the high rate of medical error, especially considering to the strain on healthcare systems by the booming elderly population, the rise of genomics and personalized medicine, their impact on the pharmaceutical industry, belief systems and ethics, and their impact on the increased need for management of privacy and consent.

Paul Peters
Applied Futurology
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of…
Paul Oude Luttighuis
Adviseur bij Le Blanc Advies
Kenneth Lloyd
Software for Modeling, Simulation and Visualization
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of…
Reply
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of…
Charles Rosenbury
Designer, Architect, Philosopher
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of…
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of Data…
Allen Woods
Advanced Meta Data – Rule Based Artificial Intelligence at THE PERFORMANCE ORGANISERS LIMITED
Allen Woods
Advanced Meta Data – Rule Based Artificial Intelligence at THE PERFORMANCE ORGANISERS LIMITED
Allen Woods
Advanced Meta Data – Rule Based Artificial Intelligence at THE PERFORMANCE ORGANISERS LIMITED
Srinidhi Boray
Founder at Ingine Inc., Democratization of locked up Medical Knowledge and Commoditization of Data…
John Coghlan
Technical writing, process mapping and improvement